upgma#
- biotite.sequence.phylo.upgma(distances)[source]#
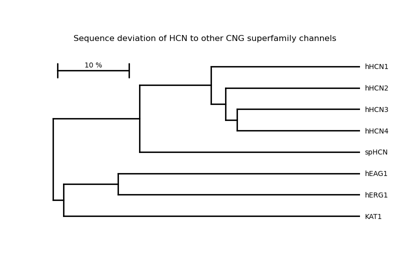
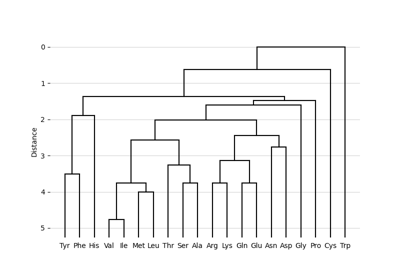
Perform hierarchical clustering using the unweighted pair group method with arithmetic mean (UPGMA).
This algorithm produces leaf nodes with the same distance to the root node. In the context of evolution this means a constant evolution rate (molecular clock).
- Parameters:
- distancesndarray, shape=(n,n)
Pairwise distance matrix.
- Returns:
- treeTree
A rooted binary tree. The index attribute in the leaf
TreeNodeobjects refer to the indices of distances.
- Raises:
- ValueError
If the distance matrix is not symmetric or if any matrix entry is below 0.
Examples
>>> distances = np.array([ ... [0, 1, 7, 7, 9], ... [1, 0, 7, 6, 8], ... [7, 7, 0, 2, 4], ... [7, 6, 2, 0, 3], ... [9, 8, 4, 3, 0], ... ]) >>> tree = upgma(distances) >>> print(tree.to_newick(include_distance=False)) ((4,(3,2)),(1,0));