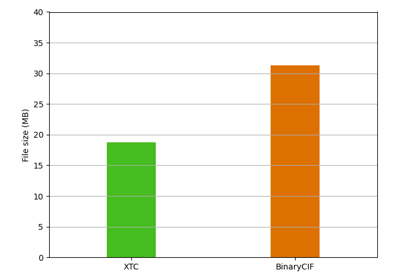
biotite.structure.io.pdbx.BinaryCIFFile¶
- class biotite.structure.io.pdbx.BinaryCIFFile(blocks=None)[source]¶
Bases:
File,_HierarchicalContainerThis class represents a BinaryCIF file.
The categories of the file can be accessed and modified like a dictionary. The values are
BinaryCIFBlockobjects.To parse or write a structure from/to a
BinaryCIFFileobject, use the high-levelget_structure()orset_structure()function respectively.Notes
The content of BinaryCIF files are lazily deserialized: Only when a column is accessed, the time consuming data decoding is performed. The decoded
BinaryCIFBlock/BinaryCIFCategoryobjects are cached for subsequent accesses.Examples
Read a BinaryCIF file and access its content:
>>> import os.path >>> file = BinaryCIFFile.read(os.path.join(path_to_structures, "1l2y.bcif")) >>> print(file["1L2Y"]["citation_author"]["name"].as_array()) ['Neidigh, J.W.' 'Fesinmeyer, R.M.' 'Andersen, N.H.'] >>> # Access the only block in the file >>> print(file.block["entity"]["pdbx_description"].as_item()) TC5b
Create a BinaryCIF file and write it to disk:
>>> category = BinaryCIFCategory({"some_column": "some_value"}) >>> block = BinaryCIFBlock({"some_category": category}) >>> file = BinaryCIFFile({"some_block": block}) >>> file.write(os.path.join(path_to_directory, "some_file.bcif"))
- Attributes
- blockBinaryCIFBlock
The sole block of the file. If the file contains multiple blocks, an exception is raised.
- clear() None. Remove all items from D.¶
- copy()¶
Create a deep copy of this object.
- Returns
- copy
A copy of this object.
- static deserialize(content)¶
Create this component by deserializing the given content.
- Parameters
- contentstr or dict
The content to be deserialized. The type of this parameter depends on the file format. In case of CIF files, this is the text of the lines that represent this component. In case of BinaryCIF files, this is a dictionary parsed from the MessagePack data.
- get(k[, d]) D[k] if k in D, else d. d defaults to None.¶
- items() a set-like object providing a view on D's items¶
- keys() a set-like object providing a view on D's keys¶
- pop(k[, d]) v, remove specified key and return the corresponding value.¶
If key is not found, d is returned if given, otherwise KeyError is raised.
- popitem() (k, v), remove and return some (key, value) pair¶
as a 2-tuple; but raise KeyError if D is empty.
- classmethod read(file)¶
Read a BinaryCIF file.
- Parameters
- filefile-like object or str
The file to be read. Alternatively a file path can be supplied.
- Returns
- file_objectBinaryCIFFile
The parsed file.
- serialize()¶
Convert this component into a Python object that can be written to a file.
- Returns
- contentstr or dict
The content to be serialized. The type of this return value depends on the file format. In case of CIF files, this is the text of the lines that represent this component. In case of BinaryCIF files, this is a dictionary that can be encoded into MessagePack.
- setdefault(k[, d]) D.get(k,d), also set D[k]=d if k not in D¶
- static subcomponent_class()¶
Get the class of the components that are stored in this component.
- Returns
- subcomponent_classtype
The class of the subcomponent. If this component already represents the lowest level, i.e. it does not contain subcomponents,
Noneis returned.
- static supercomponent_class()¶
Get the class of the component that contains this component.
- Returns
- supercomponent_classtype
The class of the supercomponent. If this component present already the highest level, i.e. it is not contained in another component,
Noneis returned.
- update([E, ]**F) None. Update D from mapping/iterable E and F.¶
If E present and has a .keys() method, does: for k in E: D[k] = E[k] If E present and lacks .keys() method, does: for (k, v) in E: D[k] = v In either case, this is followed by: for k, v in F.items(): D[k] = v
- values() an object providing a view on D's values¶
- write(file)¶
Write contents into a BinaryCIF file.
- Parameters
- filefile-like object or str
The file to be written to. Alternatively, a file path can be supplied.
Gallery¶

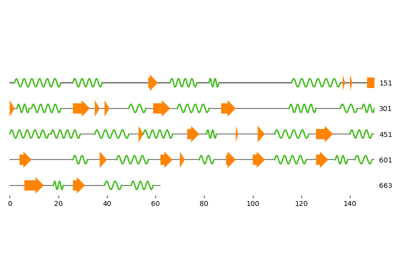
Visualization of normal modes from an elastic network model
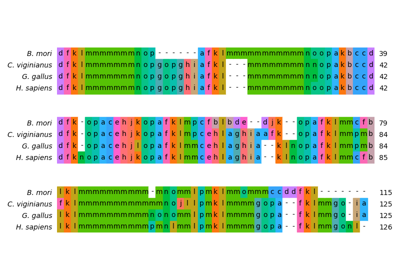
Structural alignment of lysozyme variants using ‘Protein Blocks’
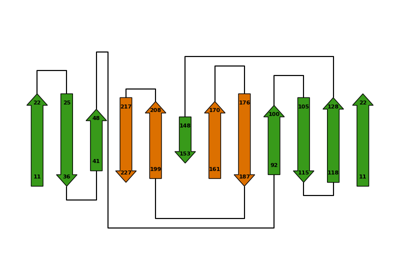
Three ways to get the secondary structure of a protein