align_multiple#
- biotite.sequence.align.align_multiple(sequences, matrix, gap_penalty=-10, terminal_penalty=True, distances=None, guide_tree=None)[source]#
Perform a multiple sequence alignment using a progressive alignment algorithm. [1]
Based on pairwise sequence distances a guide tree is constructed. The sequences are progessively aligned according to the tree, following the rule ‘Once a gap, always a gap’.
- Parameters:
- sequenceslist of Sequence
The sequences to be aligned. The alpahbet of the substitution matrix must be equal or extend the alphabet of each sequence.
- matrixSubstitutionMatrix
The substitution matrix used for scoring. Must be symmetric.
- gap_penaltyint or tuple(int, int), optional
If an integer is provided, the value will be interpreted as general gap penalty. If a tuple is provided, an affine gap penalty is used. The first integer in the tuple is the gap opening penalty, the second integer is the gap extension penalty. The values need to be negative.
- terminal_penaltybool, optional
If true, gap penalties are applied to terminal gaps.
- distancesndarray, shape=(n,n)
Pairwise distances of the sequences. The matrix must be symmetric and all entries must be larger than 0. By default the pairwise distances are calculated from similarities obtained from optimal global pairwise alignments (
align_optimal()). The similarities are converted into distances using the method proposed by Feng & Doolittle [2].- guide_treeTree
The guide tree to be used for the progressive alignment. By default the guide tree is constructed from distances via the UPGMA clustering method.
- Returns:
- alignmentAlignment
The global multiple sequence alignment of the input sequences.
- orderndarray, dtype=int
The sequence order represented by the guide tree. When this order is applied to alignment sequence order, similar sequences are adjacent to each other.
- treeTree
The guide tree used for progressive alignment. Equal to guide_tree if provided.
- distance_matrixndarray, shape=(n,n), dtype=float32
The pairwise distance matrix used to construct the guide tree. Equal to distances if provided.
Notes
The similarity to distance conversion is performed according to the following formula:
\[D_{a,b} = -\ln\left( \frac { S_{a,b} - S_{a,b}^{rand} } { S_{a,b}^{max} - S_{a,b}^{rand} } \right)\]\[S_{a,b}^{max} = \frac{ S_{a,a} + S_{b,b} }{ 2 }\]\[S_{a,b}^{rand} = \frac{1}{L_{a,b}} \left( \sum_{x \in \Omega} \sum_{y \in \Omega} s_{x,y} \cdot N_a(x) \cdot N_b(y) \right) + N_{a,b}^{open} \cdot p^{open} + N_{a,b}^{ext} \cdot p^{ext}\]\(D_{a,b}\) - The distance between the sequences a and b.
\(S_{a,b}\) - The similarity score between the sequences a and b.
\(s_{x,y}\) - The similarity score between the symbols x and y.
\(\Omega\) - The sequence alphabet.
\(N_a(x)\) - Number of occurences of symbol x in sequence a.
\(N_{a,b}^{open}, N_{a,b}^{ext}\) - Number of gap openings/ extensions, in the alignment of a and b.
\(p^{open}, p^{ext}\) - The penalty for a gap opening/extension.
\(L_{a,b}\) - Number of columns in the alignment of a and b.
In rare cases of extremely unrelated sequences, \(S_{a,b}\) can be lower than \(S_{a,b}^{rand}\). In this case the logarithm cannot be calculated and a
ValueErroris raised.References
Examples
>>> seq1 = ProteinSequence("BIQTITE") >>> seq2 = ProteinSequence("TITANITE") >>> seq3 = ProteinSequence("BISMITE") >>> seq4 = ProteinSequence("IQLITE") >>> matrix = SubstitutionMatrix.std_protein_matrix() >>> >>> alignment, order, tree, distances = align_multiple( ... [seq1, seq2, seq3, seq4], matrix ... ) >>> >>> print(alignment) BIQT-ITE TITANITE BISM-ITE -IQL-ITE >>> print(alignment[:, order.tolist()]) -IQL-ITE BISM-ITE BIQT-ITE TITANITE >>> print(distances) [[0.000 1.034 0.382 0.560] [1.034 0.000 0.923 1.132] [0.382 0.923 0.000 0.632] [0.560 1.132 0.632 0.000]] >>> >>> print(tree.to_newick( ... labels=["seq1", "seq2", "seq3", "seq4"], include_distance=False ... )) ((seq4,(seq3,seq1)),seq2);
Gallery#
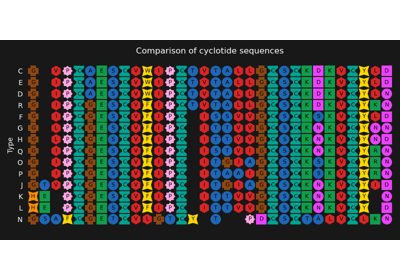
Customized visualization of a multiple sequence alignment
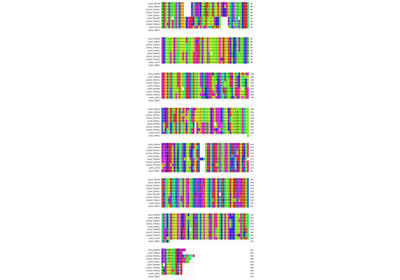

Fetching and aligning a protein from different species

Multiple Structural alignment of orthologous proteins
