biotite.sequence.io.genbank.get_annotated_sequence¶
- biotite.sequence.io.genbank.get_annotated_sequence(gb_file, format='gb', include_only=None)[source]¶
Get an annotated sequence by combining the ANNOTATION and ORIGIN fields of a GenBank file.
- Parameters
- gb_fileGenBankFile
The GenBank file to read the fields from.
- include_onlyiterable object of str, optional
List of names of feature keys, which should included in the annotation. By default all features are included.
- Returns
- annot_seqAnnotatedSequence
The annotated sequence.
Gallery¶
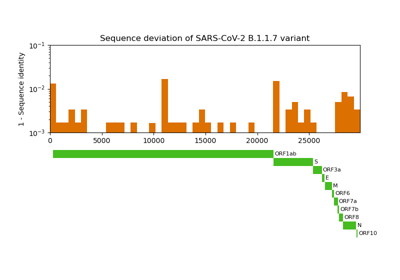
Comparative genome assembly of SARS-CoV-2 B.1.1.7 variant
Comparative genome assembly of SARS-CoV-2 B.1.1.7 variant