Biotite documentation#
Your multitool for bioinformatics
Whether you want to find homologous sequence regions in a protein family or would like to identify secondary structure elements in a protein structure: Biotite has the right tool for you. This package bundles popular tasks in computational molecular biology into a uniform and fast Python library.
Skip writing code for basic functionality and focus on what your code makes unique - from small analysis scripts to entire bioinformatics software packages.
Features
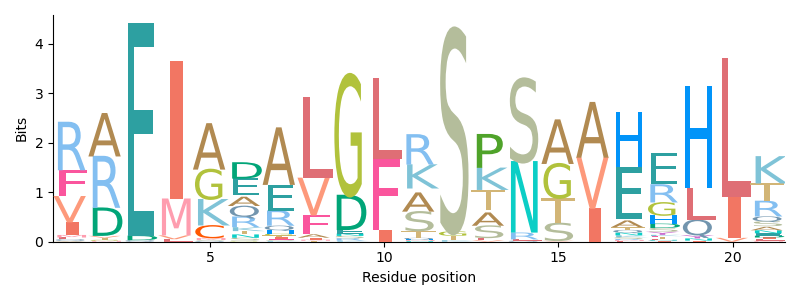
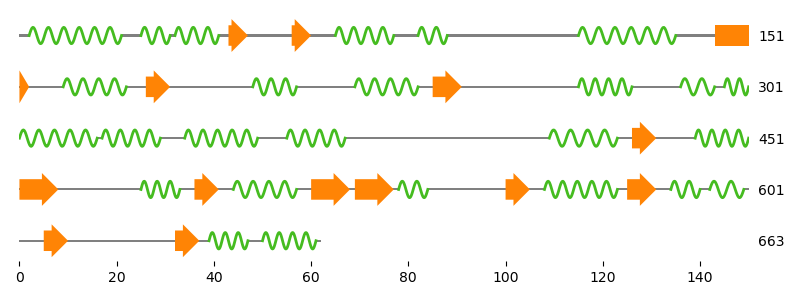
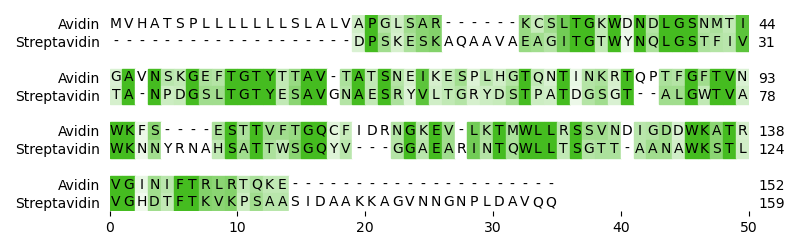
Analyze sequence data
Work with sequences of any kind: from the usual nucleotide and protein sequences to structural alphabets like 3Di. Use the rapid and modular alignment tools to identify homologous regions or to map reads. Eventually, visualize your results in different Matplotlib based representations, ranging from sequence alignments to feature maps.
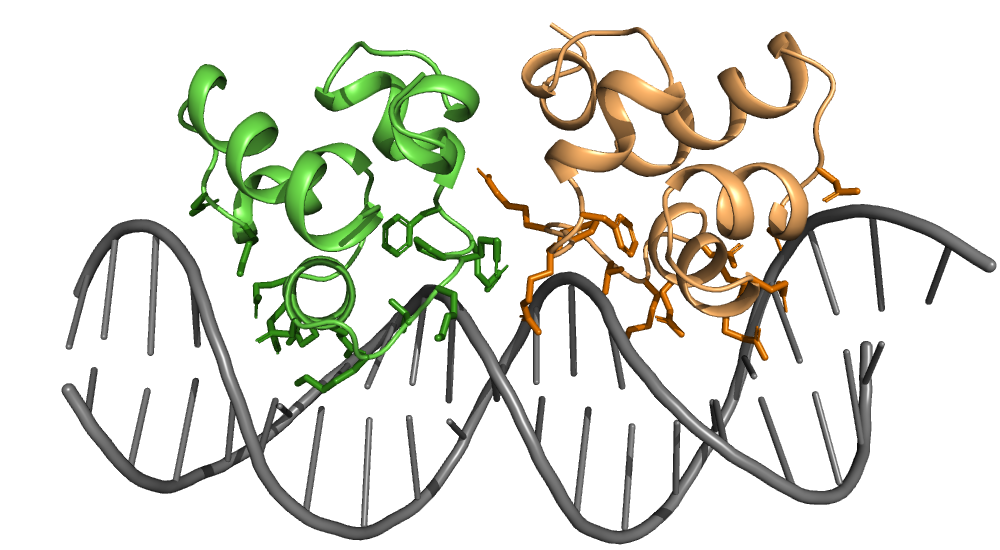
Explore molecular 3D structures
Handle 3D structures of large biomolecules as well as small molecules in an intuitive NumPy-like way. Explore a variety of functions to filter, transform and analyze your structure data - from surface area calculation to structure superimposition. Interfaces to a multitude of popular file formats such as PDB, CIF/BinaryCIF, MOL/SDF, trajectory formats and more are available.
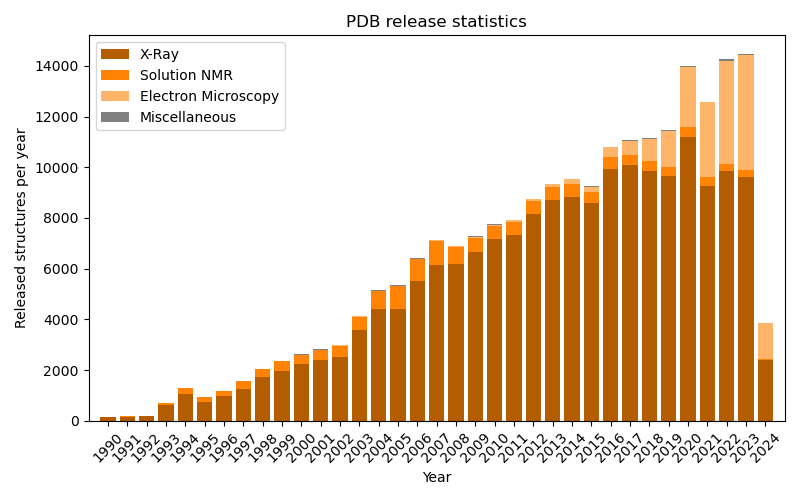
Access data in biological databases
Obtain data from biological sequence and structure databases, such as NCBI Entrez, UniProt, PDB and PubChem. Craft your queries with the help of Pythonic logical operators instead of learning the respective REST API.
Integrate popular software seamlessly
In case Biotite’s integrated functionality is not sufficient for your tasks, you can use interfaces to prominent external software ranging from multiple sequence alignment to secondary structure annotation tools. These interfaces are seamless: You can input Python objects and you get Python objects. File creation and command line execution are handled under the hood.
Support
Citation
If you use Biotite in a scientific publication, please cite one of the following articles:
P. Kunzmann, K. Hamacher, "Biotite: a unifying open source computational biology framework in Python," BMC Bioinformatics, vol. 19, pp. 346, October 2018. doi: 10.1186/s12859-018-2367-z
P. Kunzmann, T. D. Müller, M. Greil, J. H. Krumbach, J. M. Anter, D. Bauer, F. Islam, K. Hamacher, "Biotite: new tools for a versatile Python bioinformatics library," BMC Bioinformatics, vol. 24, pp. 1--19, June 2023. doi: 10.1186/s12859-023-05345-6